Get Data
Summary:
This model product contains the source code for the Ecosystem Demography Model (ED version 1.0) as well as model input and output data for a portion of South America including the Brazilian Amazon. The model output data are estimates of potential average live biomass (kg C/m2), potential average soil carbon (kg C/m2), and potential above-ground net primary production (NPP) (kg C/m2/yr) at 1.0 degree resolution. To produce these estimates, ED was forced with ISLSCP I data for 1987 and 1988, averaged into a single year (Moorcroft et al., 2001). Data for the three estimates are provided in both ASCII text and in NetCDF formatted files.
ED is an individual-based terrestrial ecosystem model that predicts both ecosystem structure (e.g. above and below-ground biomass, vegetation height and basal area, and soil carbon stocks) and corresponding ecosystem fluxes (e.g. NPP, NEP and evapotranspiration) from climate, soil, and land-use inputs. The model consists of integrated sub-models governing processes such as leaf-level physiology, plant allocation, allometry, phenology, dispersal, the effects of fire disturbances, and below-ground sub-models for soil carbon dynamics and hydrology. Using a new method for scaling-up it is possible to predict ED's large-scale behavior without simulating the fate of every plant individually. ED is used to examine how climate and edaphic factors, natural disturbances, and human land-use practices affect ecosystem structure and fluxes.
This data set contains six zip files which each uncompress into six unique subdirectories. Each subdirectory is described in detail in the Model Product Description section of this document. Installation and execution instructions are provided in the Model Documentation and User's Guide section of this document.
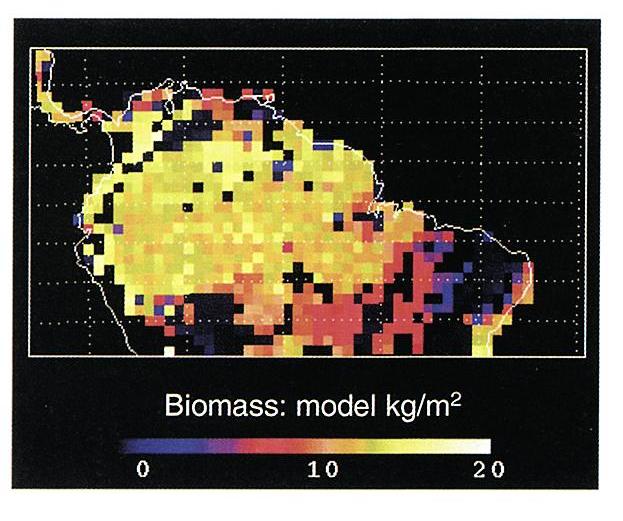
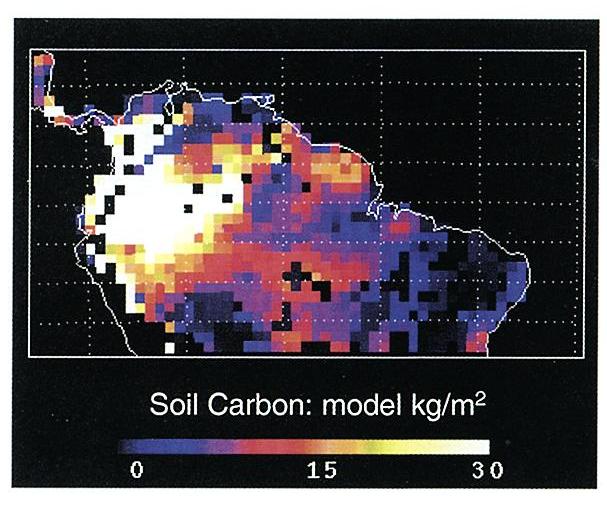
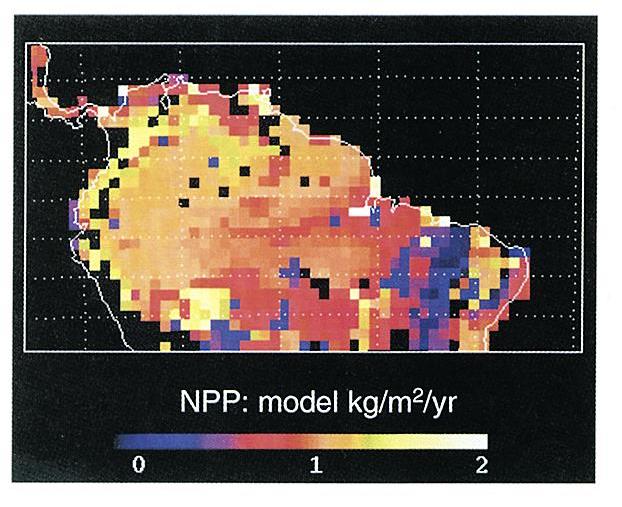
Figure 1. Model predictions for the Brazilian Amazon at
1.0 degree resolution of average above and below-ground live biomass
(kg C/m2), soil carbon stocks (kg C/m2), and average annual
above-ground NPP (kg C/m2/yr) ( Moorcroft et al., 2001).
Data Citation:
Cite this data model as follows:
Hurtt, G.C., P.R. Moorcroft, and S.W. Pacala. 2013. Ecosystem Demography Model: Scaling Vegetation Dynamics Across South America. ORNL DAAC, Oak Ridge, Tennessee, USA. http://dx.doi.org/10.3334/ORNLDAAC/1149
Links to Related Data Products:
Ecosystem Demography Model: U.S. Ecosystem Carbon Stocks and Fluxes, 1700-1990
LBA-ECO LC-08 Ecosystem Demography Model Estimated C, NPP, and Biomass for Amazonia [http://daac.ornl.gov/cgi-bin/dsviewer.pl?ds_id=1102]
Model Product Description
Data Set Files:
| Data Files | Description |
|---|---|
| code.zip | This compressed file, code.zip, expands to a sub-directory: /code/ containing the ED model code files for South America, written in C. |
| cvode.zip | This compressed file, cvode.zip, expands to a sub-directory: /cvode/ containing the ED model integration code, written in C. |
| ic.zip | This compressed file, ic.zip, expands to an empty sub-directory: /ic/ that is required for the ED model to run. |
| output.zip | This compressed file, output.zip, expands to an empty sub-directory: /output/ that is required for the ED model to run. |
| inputs.zip | This compressed file, inputs.zip, expands to the following sub-directories: /grid_cell_area/; /mask/; /mechanism.in/; /precip/; /soils/; /soil_temp/; /sois/; and /temp/. Please note that the /mechanism.in subdirectory contains two sub-directories, C3 and C4. See Input data section for complete file descriptions. |
| published_output.zip | This compressed file, published_output.zip, expands to a sub-directory: /published_output/ containing three NetCDF (.nc) and three ASCII text files (.txt) described below in the published output data section. |
Model Code Files:
The structure of this data set as it extrapolates from the zip files has been designed to ensure simplistic model installation and execution. The following zip files contain either model code or required subdirectory structures: code.zip, cvode.zip, ic.zip, and output.zip. Please see the "Model Documentation and User's Guide" section below for instructions on how to run ED.
Input Data:
There are four types of input data: Grid Cell Area; Land Sea Mask; Mechanism Input; and ISLSCP I Initiative data. These extrapolate into 8 subdirectories (grid_cell_area, mask, mechanism, precip, soils, soil_temp; sois, and temp) each described as follows:
Grid Cell Area. The grid_cell_area subdirectory contains a single file, GRIDAREA.GRD, which is a one degree global ASCII map of grid cell area provided in km2, and provided by Moorcroft et al. 2001.
Mask. The mask subdirectory contains a single file, land_sea.msk, which is a land/sea mask where a value of 1 represents land and a value of 0 represents water. This file is in ascii text format at a spatial resolution of 1 degree latitude by 1 degree longitude arranged in 360 columns and 180 rows.
Mechanism Input. The files in the mechanism subdirecory (lat**long**.in) provide pre-computed, monthly-integrated photosynthesis and transpiration per leaf area for 121 light levels within the canopy. There are two sets of files, one for each physiology (C3 and C4) with one file per grid cell. Each file contains 5 columns for each light level for each month and includes: fraction of full light; photosynthesis with stomates open (g C/m2/month); photosynthesis with stomates closed (g C/m2/month); transpiration with stomates open (g Water/m2/month); and transpiration with stomates closed (g Water/m2/month).
ISLSCP I Initiative Data. For the ED South American data set, the model was run using data from the International Satellite Land Surface Climatology Project (ISLSCP) Initiative I. These data input files provide modelers with many of the fields required to prescribe boundary conditions and initialize and force a wide range of land-biosphere-atmosphere models. All of these data have been preprocssed to the same spatial resolution (1 degree by 1 degree), using the same land/sea mask and processing steps to ensure spatial and temporal continuity. These data cover the period 1987 through 1988 at a 1-month time resolution for most of the seasonally varying quantities and at 6-hourly resolution for the near-surface meteorological and radiative forcings. The data for 1987 and 1988 were averaged into a single average year. Input data from ISLSCP Iniative I include the following:
| SUB-DIRECTORY | FILE NAMES | VARIABLES | UNITS | SOURCE |
|---|---|---|---|---|
| precip | precip.islscp.amaz | Monthly mean precipitation amount (based on gauge measurements) | mm | GPCP/GPCC |
| soil_temp | soil_temp.islscp.amaz | Monthly mean depth soil temperature | Kelvin | ECMWF |
| soils | amaz.soils.dat | Soil depth (estimate of the depth from the soil surface to bedrock or other impermeable layer) | mm | NASA: Webb et al. (1993) |
| Soil texture (classification: 1=coarse; 2=medium/coarse; 3=medium; 4=fine/medium; 5=fine; 6=ice; 7=organic; 0=ocean) | Index (0-7) | NASA: Zobler et al. (1986) | ||
| Theta_max (matrix potential at saturation) | m | Meeson et al. (1995); Sellers et al. (1995) | ||
| k_sat (saturated hydraulic conductivity or Ksat) | m/s | |||
| tau (an empirical power governing the rate at which conductivity decreases as saturation levels decrease) | dimensionless | |||
| temp | temp.islscp.amaz | Monthly average air temperature (6-hourly resolution) at 2 meters above the ground | degrees Celsius | ECMWF |
| sois | soi.list | Locations of South American sites that were used to evaluate the size- and age-structured (SAS) approximation | decimal degrees | Moorcroft et al. (2001) |
| mechanism.in |
Contains folders c3 and c4. Example file
name contained within the folders: lat25lon-99.in |
The mechanism files (lat**lon**.in) provide pre-computed, monthly-integrated photosynthesis and transpiration per leaf area for 121 light levels within the canopy. There are two sets of files, one for each physiology (C3 and C4) with one file per grid cell. | Files list 5 columns for each light level for each month:
|
Moorcroft et al. (2001) |
| grid_cell_area | GRIDAREA.GRD | One degree global ASCII map of grid cell area | km2 | Moorcroft et al. (2001) |
| mask | land_sea.msk | Land - sea mask | Value of 1 for land and 0 for water | |
|
Notes: GPCP/GPCC = Global Precipitation Climatology Project /Global Precipitation Climatology Center. |
||||
| ECMWF = European Center for Medium-Range Weather Forecasts. | ||||
| Climate and soil properties within each 1 degree grid cell are derived from the ISLSCP (International Satellite Land Surface Climatology Project) Initiative I global data set (Meeson et al., 1995; Sellers et al., 1995). | ||||
| The ISLSCP data set was specifically designed for large-scale biosphere modeling, providing a consistent global data set of climatological variables and soil characteristics. | ||||
| A preliminary review suggested that the data set is comparable in quality to other global data sets (Kerr, 1995); however, subsequent analyses have identified several shortcomings. In particular, while the data set provides global coverage, its temporal coverage is short, containing only two representative years of climate data. | ||||
| With regard to the South American continent, the quality of the precipitation data is reduced because of sparse gauge coverage (Kerr, 1995), and the estimate of downward shortwave radiation over the Amazon basin is thought to contain a significant degree of error (Morrill, 1999). | ||||
| Although the implementation described here relies on ISLSCP data, the model can be driven from other sources of data including output from a climate model. Source: Moorcroft et al. (2001). | ||||
Published Output Data:
The ED output data files for South America (Amazonia) (Moorcroft et al., 2001) are provided in both ASCII tabular (.txt) and NetCDF (.nc) (Network Common Data Format) format.
| FILE NAME | VARIABLES | UNITS | DATA RANGE |
|---|---|---|---|
| ED_SA_v1_Soil.nc | Potential Soil Carbon | kg C/m2 | 0.000043 – 29.88 |
| ED_SA_v1_Soil.txt | |||
| ED_SA_v1_Biomass.nc | Potential Live Biomass | kg C/m2 | 0 - 18.61 |
| ED_SA_v1_Biomass.txt | |||
| ED_SA_v1_NPP.nc | Potential Net Primary Production | kg C/m2/yr (monthly) | - 0.57 – 4.54 |
| ED_SA_v1_NPP.txt |
Spatial & Temporal Scales
The Amazon data set has a grid cell size of 1 degree by 1 degree. This data set covers southern Latin America and the northern half of South America from 15 degrees N to 15 degrees S and spans from 30 degrees W to 85 degrees W longitude. The data represent a snapshot the time period covered by the input variables (1987/01/01-1988/12/31) averaged into a single average year.
Data Format
The output data files are in ASCII and NetCDF formats. The full data set contains 30 rows [the x-dimension (longitude)] by 55 columns [the y-dimension (latitude)] in the the full spatial extent. Data are 32-bit floating point data. The fill value for missing output data is –9.9. The ISLSCP Initiative I and C3 and C4 mechanism input data are ASCII text files. The fill value for missing input data values is -99. The fill value for non-land/ocean grid cells (sea mask) is 0.
Variable Description:
a) Potential Soil Carbon
Ecosystem Demography Model (ED) estimates of potential average soil carbon (kg C/m2), including litter and necromass, at 1 degree resolution. These estimates include the effects of natural disturbances such as windthrow and fire, but do not include the effects of human land use.
b) Potential Live Biomass
ED estimates of potential average live biomass (kg C/m2) at 1 degree resolution. These estimates include the effects of natural disturbances such as windthrow and fire, but do not include the effects of human land use.
c) Potential Net Primary Production
ED estimates the potential net primary production per grid cell (kg C/m2/yr), on a monthly basis. These estimates include the effects of natural disturbances such as windthrow and fire, but do not include the effects of human land use.
Example Data Record from ED_SA_v1_Biomass.txt
| Band Name: biomass Description: 1 degree grid cell size. Ecosystem Demography Model (ED) for Amazonia: Estimate of potential live above and below ground biomass. See data references and details in the Auxiliary file or the Data Guide. Holding name: Potential Live Biomass Collection name: Ecosystem Demography Model Data Date class: snapshot Begin date of output End date of output Time step units: snapshot Data units: kg C m-2 Fill value: -9.9 Center Lon,Center Lat,snapshot -84.500000,14.500000,13.210 -83.500000,14.500000,12.413 -82.500000,14.500000,-9.900 -81.500000,14.500000,-9.900 ... -33.500000,-14.500000,-9.900 -32.500000,-14.500000,-9.900 -31.500000,-14.500000,-9.900 -30.500000,-14.500000,-9.900 |
Note: Natural disturbances and human land-use activities have a large impact on the structure, dynamics and fluxes of ecosystems at a variety of scales. As a result, ecosystems are more heterogeneous in important ways than is explained by climatic and edaphic factors alone. A critical impediment to incorporating relevant fine-scale detail in large-scale models is the problem of scaling-up.
Site boundaries: (All latitude and longitude given in decimal degrees)
| Site (Region) | Westernmost Longitude | Easternmost Longitude | Northernmost Latitude | Southernmost Latitude | Geodetic Datum |
|---|---|---|---|---|---|
| South America (including Amazonia) | -85 | -30 | 15 | -15 | World Geodetic System, 1984 (WGS-84) |
Model Documentation and User's Guide
Description:
The ED model solves a system of size- and age-structured partial differential equations (PDEs) for each grid cell numerically using the method of characteristics. The model, written in C, makes extensive use of C structures, and pointers and dynamic memory allocation.
Steps for compiling and running the Ecosystem Demography Model (ED):
1. Build the integration library:
The ED model uses Scott D. Cohen's and Alan C. Hindmarsh's CVODE
integration
code. To build the integration library, cd to the /cvode/ subdirectory.
Check
the Makefile to ensure the C compiler, paths and compile flags are
appropriate
for your system. Compile the library by typing 'make'. If successful,
the
library libcvode.a will appear in /cvode/lib. Then cd to /cvode/lib and
type
'ranlib libcvode.a'. The library will now be ready to use.
2. Compile the ED model executable:
cd to the /code sub-directory of the ED directory tree. Edit the
Makefile to
set the C compiler, paths and compile flags appropriately for your
system.
Edit the header file ed.h, using the macros to set desired options.
Details of the different options are included in the header file comments.
Important settings include:
TMAX - float indicating the number of years to be integrated (eg 200.0)
REGION - integer flag. Set to 0 to run model for the grid cells
specified in
inputs/sois/soi.list. Set to 1 perform a regional integration.
RESTART - integer flag indicating whether run is a restart. Generally
set to 0.
This begins integration from a low initial density (0.1 indivs/m2) of
seedlings
(size HGTMIN) of each plant functional type . When RESTART is set to 1,
initial
conditions are read in from restart files in the ic/ subdirectory.
Before
performing a restart, restart files from the previous integration
period should
be moved or copied to the ic/ subdirectory. See output section below
for more
details on restart files.
Build the ED model executable (edm) then by typing 'make'.
Additional model parameters are specified the init_data sub-routine
found in
init_data.c source file
3. Run ED:
Run the ED model in by typing 'edm'. Output indicating the progress of
the
integration will be printed to the stdout and output data will
accumulate in
the output/ subdirectory (see below for details). The frequency of
printing is
set by the value of the macro PRINT_OUTPUT_FREQ.
Output Files
During integration, data is output into files in the output/
subdirectory.
For the grid-cells specified in inputs/sois/soi.list, numerous files of
the
form pfx.latxxlonyy.* (where pfx is the prefix specified in the ed.h
header and
xx and yy are the lat/lon coordinates of the North-West corner the grid
cell).
These files contain detailed information on the ecosystem structure and
fluxes
for the grid cell. Files with the word 'patch' contain contain
information on
the age-related subgrid-scale variability in ecosystem structure and
fluxes.
Consult model source files for more details on the various quantities
output.
If a regional integration is performed, the model will also output
numerous files of the form pfx.region.* These files contain gridded region-wide
snapshots of model quantities. Output is in the form of a [lon] x [lat]
x [time] matrix of values. Longitude varies fastest followed by latitude,
then time. Consult model source files for more details on the quantities
output.
During the integration two restart files per grid cell (pfx.latxxlonyy.pss and
pfx.latxxlonyy.css) are created periodically (frequency is determined by the
value of the macro PRINT_RESTART_FREQ).
Data Access:
This data is available through the Oak Ridge National Laboratory (ORNL) Distributed Active Archive Center (DAAC).
Data Archive Center:
Contact for Data Center Access Information:
E-mail: uso@daac.ornl.gov
Telephone: +1 (865) 241-3952
References:
Moorcroft, P.R, G.C. Hurtt, and S.W. Pacala. 2001. A method for scaling vegetation dynamics: the ecosystem demography model (ED). Ecological Monographs 71: 557-585.
Moorcroft, P.R. G.C. Hurtt, and S.W. Pacala. 2001. A method for scaling vegetation dynamics: the ecosystem demography model (ED). Appendices. Ecological Archives M071-008.
Additional Sources of Information:
Hurtt G.C., P.R. Moorcroft, S.W. Pacala, and S.A. Levin. 1998. Terrestrial models and global change: challenges for the future. Global Change Biology 4: 581-590.
Kerr, Y.H. 1995. A review of the ISLSCP Initiative I CD-ROM collection: context, scope, and main outcome. Goddard Space Flight Center, National Aeronautics and Space Administration, Greenbelt, Maryland, USA.
Meeson, B W., F.E. Corprew, J.M.P. McManus, D.M. Myers, J.W. Closs, K.J. Sun, D J. Sunday, and P.J. Sellers. 1995. ISLSCP Initiative I-global data sets for land-atmosphere models, 1987-1988. American Geophysical Union, Washington, D.C., U.S.A. [Available on CD-ROM.]
Morrill, J.C. 1999. Sensitivity of a land surface model to the diurnal distribution of downward longwave radiation. Journal of the Meteorological Society of Japan 77: 265-279.
Sellers, P.J., et al. 1995. An overview of the ISLSCP Initiative I global data sets. In Meeson, B W., F.E. Corprew, J.M.P. McManus, D.M. Myers, J.W. Closs, K.J. Sun, D J. Sunday, and P.J. Sellers. ISLSCP Initiative I-global data sets for land-atmosphere models, 1987-1988. American Geophysical Union, Washington, D.C., USA. [Available on CD-ROM.]
Webb, R.S., C.E. Rosenzweig, and E.R. Levine. 1993. Specifying land surface characteristics in general circulation models: soil profile data set and derived water-holding capacities. Global Biogeochemical Cycles 7: 97-108.
Zobler, L. 1986. A world soil file for global climate modeling. NASA Tech. Memo. 8780. NASA pp. 36.